Note
Go to the end to download the full example code.
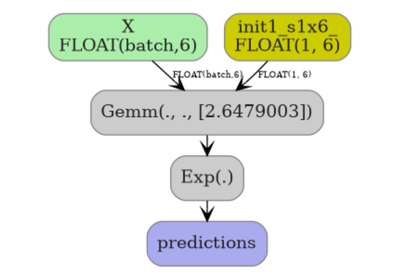
KNeighbors: choosing between CDist and standard ONNX#
yobx.sklearn.to_onnx() converts a fitted
sklearn.neighbors.KNeighborsClassifier or
sklearn.neighbors.KNeighborsRegressor into an
onnx.ModelProto.
Two implementations are available, selected automatically at conversion time
based on the target_opset argument:
Standard ONNX (default) — uses built-in ONNX operators only (
Mul,ReduceSum,MatMul,TopK, …). Runs on any ONNX-compatible runtime. Requires at least opset 13 for the classifier (ReduceSumwith axes-as-input) and opset 18 for the regressor (ReduceMeanwith axes-as-input).CDist path (opt-in) — delegates pairwise distance computation to the
com.microsoft.CDistcustom operator, which is natively accelerated by ONNX Runtime. Enable it by passingtarget_opset={"": <n>, "com.microsoft": 1}.
When to prefer CDist#
Use the CDist path when all of the following are true:
You are deploying to ONNX Runtime (which ships with CDist).
The training set is large (many rows M in
_fit_X). CDist is fused at the C++ level and avoids materialising the full(N, M)intermediate matrix in Python.You are comfortable with a model that cannot run on runtimes that do not implement the
com.microsoftcustom domain.
Use the standard ONNX path when you need maximum portability.
import numpy as np
import onnxruntime
from sklearn.neighbors import KNeighborsClassifier, KNeighborsRegressor
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from yobx.sklearn import to_onnx
1. Train a KNeighborsClassifier#
rng = np.random.default_rng(0)
X = rng.standard_normal((80, 4)).astype(np.float32)
y = (X[:, 0] > 0).astype(np.int64)
clf = KNeighborsClassifier(n_neighbors=5)
clf.fit(X, y)
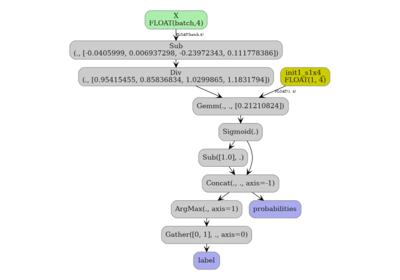
Standard ONNX path (default)#
Pass a plain integer opset (or omit target_opset entirely). The
converter uses built-in ONNX operators only and the resulting model is
portable across any ONNX runtime.
onx_std = to_onnx(clf, (X,))
op_types_std = {(n.op_type, n.domain or "") for n in onx_std.graph.node}
print("Standard path nodes:", sorted({t for t, _ in op_types_std}))
assert ("CDist", "com.microsoft") not in op_types_std
assert ("TopK", "") in op_types_std
ref_std = onnxruntime.InferenceSession(
onx_std.SerializeToString(), providers=["CPUExecutionProvider"]
)
labels_std = ref_std.run(None, {"X": X})[0]
assert (labels_std == clf.predict(X).astype(np.int64)).all()
print("Standard path labels match sklearn ✓")
Standard path nodes: ['Add', 'ArgMax', 'Div', 'Gather', 'MatMul', 'Mul', 'OneHot', 'ReduceSum', 'Relu', 'Reshape', 'Sqrt', 'Sub', 'TopK']
Standard path labels match sklearn ✓
CDist path (com.microsoft)#
Pass a dict target opset that includes "com.microsoft": 1.
The converter inserts a single com.microsoft.CDist node for distance
computation, which ONNX Runtime executes via a fused C++ kernel.
onx_cd = to_onnx(clf, (X,), target_opset={"": 18, "com.microsoft": 1})
op_types_cd = {(n.op_type, n.domain or "") for n in onx_cd.graph.node}
print("CDist path nodes:", sorted({t for t, _ in op_types_cd}))
assert ("CDist", "com.microsoft") in op_types_cd
ref_cd = onnxruntime.InferenceSession(
onx_cd.SerializeToString(), providers=["CPUExecutionProvider"]
)
labels_cd = ref_cd.run(None, {"X": X})[0]
assert (labels_cd == clf.predict(X).astype(np.int64)).all()
print("CDist path labels match sklearn ✓")
CDist path nodes: ['ArgMax', 'CDist', 'Div', 'Gather', 'OneHot', 'ReduceSum', 'Relu', 'Reshape', 'TopK']
CDist path labels match sklearn ✓
Both paths produce identical predictions#
assert (labels_std == labels_cd).all(), "Paths produce different labels!"
print("Both paths agree ✓")
Both paths agree ✓
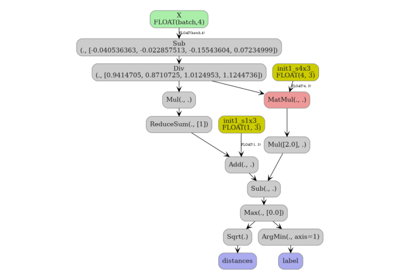
2. KNeighborsRegressor — standard ONNX path#
The regressor uses ReduceMean with axes as an input, which requires
at least opset 18.
X_r = rng.standard_normal((60, 3)).astype(np.float32)
y_r = (X_r[:, 0] * 3 + X_r[:, 1]).astype(np.float32)
reg = KNeighborsRegressor(n_neighbors=5)
reg.fit(X_r, y_r)
onx_reg = to_onnx(reg, (X_r,)) # uses opset 18 by default
ref_reg = onnxruntime.InferenceSession(
onx_reg.SerializeToString(), providers=["CPUExecutionProvider"]
)
preds_onnx = ref_reg.run(None, {"X": X_r})[0]
preds_sk = reg.predict(X_r).astype(np.float32)
assert np.allclose(preds_onnx, preds_sk, atol=1e-5), "Regressor predictions differ!"
print("Regressor predictions match sklearn ✓")
Regressor predictions match sklearn ✓
3. Inside a Pipeline#
Both implementations work transparently inside a scikit-learn
Pipeline.
pipe = Pipeline([("scaler", StandardScaler()), ("clf", KNeighborsClassifier(n_neighbors=5))])
pipe.fit(X, y)
onx_pipe = to_onnx(pipe, (X,))
ref_pipe = onnxruntime.InferenceSession(
onx_pipe.SerializeToString(), providers=["CPUExecutionProvider"]
)
labels_pipe = ref_pipe.run(None, {"X": X})[0]
assert (labels_pipe == pipe.predict(X).astype(np.int64)).all()
print("Pipeline labels match sklearn ✓")
Pipeline labels match sklearn ✓
Total running time of the script: (0 minutes 0.194 seconds)
Related examples