Note
Go to the end to download the full example code.
Exporting sklearn tree models with convert options#
yobx.sklearn.to_onnx() accepts a convert_options argument that
enables extra outputs on tree and ensemble estimators without changing
the core converter interface.
ConvertOptions exposes two boolean flags:
decision_path — an additional string tensor encoding, for each sample, the binary root-to-leaf path through the tree. Shape
(N, 1)for single trees;(N, n_estimators)for ensembles.decision_leaf — an additional
int64tensor containing the zero-based leaf node index reached by each sample. Same shape convention asdecision_path.
Both extras are appended after the standard model outputs and can be
validated against scikit-learn’s own decision_path()
and apply() methods.
import numpy as np
import onnxruntime
from sklearn.ensemble import RandomForestClassifier
from sklearn.tree import DecisionTreeClassifier
from yobx.doc import plot_dot
from yobx.sklearn import ConvertOptions, to_onnx
1. Train the models#
rng = np.random.default_rng(0)
X_train = rng.standard_normal((120, 4)).astype(np.float32)
y_train = (X_train[:, 0] + X_train[:, 1] > 0).astype(int)
X_test = rng.standard_normal((30, 4)).astype(np.float32)
dt = DecisionTreeClassifier(max_depth=4, random_state=0).fit(X_train, y_train)
rf = RandomForestClassifier(n_estimators=3, max_depth=3, random_state=0).fit(X_train, y_train)
2. decision_path for a single tree#
Passing ConvertOptions(decision_path=True) adds a third output to the
ONNX model. The extra tensor has dtype object (bytes/string) and
shape (N, 1) for a single tree. Each value is a binary string whose
i-th character is '1' when node i was visited and '0' otherwise.
opts_path = ConvertOptions(decision_path=True)
onx_dt_path = to_onnx(dt, (X_train,), convert_options=opts_path)
print("=== DecisionTreeClassifier with decision_path ===")
print(f"Model outputs: {[o.name for o in onx_dt_path.graph.output]}")
sess = onnxruntime.InferenceSession(
onx_dt_path.SerializeToString(), providers=["CPUExecutionProvider"]
)
label_onnx, proba_onnx, path_onnx = sess.run(None, {"X": X_test})
# Verify labels and probabilities against sklearn
assert np.array_equal(dt.predict(X_test), label_onnx), "Labels differ!"
assert np.allclose(
dt.predict_proba(X_test).astype(np.float32), proba_onnx, atol=1e-5
), "Probabilities differ!"
# Compare path strings to sklearn's sparse decision_path matrix
sklearn_dp = dt.decision_path(X_test).toarray()
onnx_dp = np.array([[int(c) for c in row[0]] for row in path_onnx], dtype=np.int8)
assert np.array_equal(onnx_dp, sklearn_dp), "Decision paths differ!"
print(f"path_onnx shape : {path_onnx.shape} (N={X_test.shape[0]}, 1 path per sample)")
print(f"path_onnx[0] : {path_onnx[0, 0]!r} (binary node-visit string)")
print("Labels, probabilities, and decision paths match sklearn ✓")
=== DecisionTreeClassifier with decision_path ===
Model outputs: ['label', 'probabilities', 'decision_path']
path_onnx shape : (30, 1) (N=30, 1 path per sample)
path_onnx[0] : '10000000000000101' (binary node-visit string)
Labels, probabilities, and decision paths match sklearn ✓
3. decision_leaf for a single tree#
ConvertOptions(decision_leaf=True) adds an int64 output containing
the zero-based leaf node index reached by each sample, matching
apply().
opts_leaf = ConvertOptions(decision_leaf=True)
onx_dt_leaf = to_onnx(dt, (X_train,), convert_options=opts_leaf)
print("\n=== DecisionTreeClassifier with decision_leaf ===")
print(f"Model outputs: {[o.name for o in onx_dt_leaf.graph.output]}")
sess = onnxruntime.InferenceSession(
onx_dt_leaf.SerializeToString(), providers=["CPUExecutionProvider"]
)
label_onnx, proba_onnx, leaf_onnx = sess.run(None, {"X": X_test})
expected_leaves = dt.apply(X_test).reshape(-1, 1)
assert np.array_equal(leaf_onnx, expected_leaves), "Leaf indices differ!"
print(f"leaf_onnx shape : {leaf_onnx.shape} (N={X_test.shape[0]}, 1 leaf index per sample)")
print(f"leaf_onnx[:5] : {leaf_onnx[:5, 0]}")
print("Labels, probabilities, and leaf indices match sklearn ✓")
=== DecisionTreeClassifier with decision_leaf ===
Model outputs: ['label', 'probabilities', 'decision_leaf']
leaf_onnx shape : (30, 1) (N=30, 1 leaf index per sample)
leaf_onnx[:5] : [16 3 16 3 13]
Labels, probabilities, and leaf indices match sklearn ✓
4. Ensemble models — decision_leaf#
For ensemble models the shape of the extra output becomes
(N, n_estimators), one column per tree. Here we use
RandomForestClassifier with decision_leaf
to retrieve the leaf node indices, which match
apply().
n_estimators = rf.n_estimators
opts_leaf_rf = ConvertOptions(decision_leaf=True)
onx_rf_leaf = to_onnx(rf, (X_train,), convert_options=opts_leaf_rf)
print("\n=== RandomForestClassifier with decision_leaf ===")
print(f"Model outputs: {[o.name for o in onx_rf_leaf.graph.output]}")
sess = onnxruntime.InferenceSession(
onx_rf_leaf.SerializeToString(), providers=["CPUExecutionProvider"]
)
label_onnx, proba_onnx, leaf_onnx = sess.run(None, {"X": X_test})
assert np.array_equal(rf.predict(X_test), label_onnx), "Labels differ!"
assert np.allclose(
rf.predict_proba(X_test).astype(np.float32), proba_onnx, atol=1e-4
), "Probabilities differ!"
expected_leaves_rf = rf.apply(X_test)
assert np.array_equal(leaf_onnx, expected_leaves_rf), "Ensemble leaf indices differ!"
print(f"leaf_onnx shape : {leaf_onnx.shape} (N={X_test.shape[0]}, {n_estimators} trees)")
print(f"leaf_onnx[0] : {leaf_onnx[0]}")
print("Labels, probabilities, and leaf indices match sklearn ✓")
=== RandomForestClassifier with decision_leaf ===
Model outputs: ['label', 'probabilities', 'decision_leaf']
leaf_onnx shape : (30, 3) (N=30, 3 trees)
leaf_onnx[0] : [10 10 10]
Labels, probabilities, and leaf indices match sklearn ✓
5. Ensemble models — decision_path#
decision_path=True adds binary path strings per estimator. For a
forest with k trees the output shape is (N, k).
opts_path_rf = ConvertOptions(decision_path=True)
onx_rf_path = to_onnx(rf, (X_train,), convert_options=opts_path_rf)
print("\n=== RandomForestClassifier with decision_path ===")
print(f"Model outputs: {[o.name for o in onx_rf_path.graph.output]}")
sess = onnxruntime.InferenceSession(
onx_rf_path.SerializeToString(), providers=["CPUExecutionProvider"]
)
label_onnx, proba_onnx, path_onnx = sess.run(None, {"X": X_test})
assert np.array_equal(rf.predict(X_test), label_onnx), "Labels differ!"
print(f"path_onnx shape : {path_onnx.shape} (N={X_test.shape[0]}, {n_estimators} trees)")
print(f"path_onnx[0] : {path_onnx[0]}")
print("Labels and decision paths produced ✓")
=== RandomForestClassifier with decision_path ===
Model outputs: ['label', 'probabilities', 'decision_path']
path_onnx shape : (30, 3) (N=30, 3 trees)
path_onnx[0] : ['10000000101' '10000000101' '1000000011100']
Labels and decision paths produced ✓
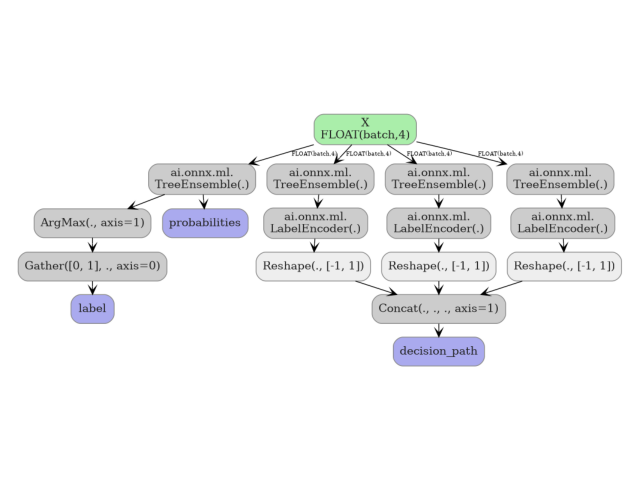
6. Visualize the model graph#
plot_dot(onx_rf_path)
Total running time of the script: (0 minutes 0.305 seconds)
Related examples