Note
Go to the end to download the full example code.
Custom converter with convert options#
This example shows how to write a custom sklearn converter whose behaviour
is controlled by a user-supplied ConvertOptionsProtocol
object.
The idea mirrors the built-in ConvertOptions
(decision_leaf, decision_path), but applied to a fully custom
estimator. The workflow has three steps:
Define the estimator — a plain scikit-learn transformer.
Define a custom options class — a lightweight object that implements the
ConvertOptionsProtocolprotocol (available_optionsandhas).Write the converter — a function that checks
g.convert_options.has("option_name", estimator)to decide whether to emit the optional extra output.
The custom estimator used here is ClipTransformer: it clips every feature
to a [clip_min, clip_max] range (equivalent to np.clip). The optional
extra output, activated by ClipOptions(clip_mask=True), is a boolean mask
tensor indicating which values were actually clipped.
import numpy as np
import onnxruntime
from sklearn.base import BaseEstimator, TransformerMixin
from yobx.doc import plot_dot
from yobx.helpers.onnx_helper import tensor_dtype_to_np_dtype
from yobx.sklearn import to_onnx
from yobx.typing import ConvertOptionsProtocol, GraphBuilderExtendedProtocol
1. Custom estimator: ClipTransformer#
A minimal transformer that clips all features into [clip_min, clip_max].
The helper method get_clip_mask returns a boolean array marking values
that were changed — it is used later to validate the ONNX output.
class ClipTransformer(TransformerMixin, BaseEstimator):
"""Clips every feature value to ``[clip_min, clip_max]``."""
def __init__(self, clip_min: float = 0.0, clip_max: float = 1.0):
self.clip_min = clip_min
self.clip_max = clip_max
def fit(self, X, y=None):
return self
def transform(self, X):
return np.clip(X, self.clip_min, self.clip_max)
def get_clip_mask(self, X):
"""Boolean mask: ``True`` where a value was clipped."""
return (X < self.clip_min) | (X > self.clip_max)
2. Custom convert options: ClipOptions#
The options class must implement two methods:
available_options()— returns the list of option names the class recognises. The framework iterates this list to decide how many extra output slots to pre-allocate for each estimator.has(option_name, piece, name=None)— returnsTruewhen the option should be active for the given fitted estimator piece. The optional name is the pipeline step name (useful for enabling an option only for a specific named step in aPipeline).
class ClipOptions(ConvertOptionsProtocol):
"""Convert options for :class:`ClipTransformer`.
:param clip_mask: when ``True``, adds a second boolean output tensor whose
value is ``True`` at each position where the input was clipped.
"""
def __init__(self, clip_mask: bool = False):
self.clip_mask = clip_mask
def available_options(self):
"""Returns the list of option names this class supports."""
return ["clip_mask"]
def has(self, option_name: str, piece: object, name=None) -> bool:
"""Returns ``True`` when *option_name* is active for *piece*."""
if option_name == "clip_mask":
# Only activate for estimators that have a clip_min attribute
# (i.e. ClipTransformer instances).
return bool(self.clip_mask) and hasattr(piece, "clip_min")
return False
3. Converter function#
The converter always emits the primary Clip output (outputs[0]).
When g.convert_options.has("clip_mask", estimator) returns True the
framework has pre-allocated outputs[1] and the converter emits
Less + Greater + Or to fill it.
def convert_clip_transformer(
g: GraphBuilderExtendedProtocol,
sts: dict,
outputs: list,
estimator: ClipTransformer,
X: str,
name: str = "clip",
) -> str:
"""Convert :class:`ClipTransformer` to ONNX.
Primary output
``outputs[0]`` — clipped values, same dtype and shape as *X*.
Optional extra output (when ``ClipOptions(clip_mask=True)`` is used)
``outputs[1]`` — boolean mask, ``True`` where a value was clipped.
"""
itype = g.get_type(X)
dtype = tensor_dtype_to_np_dtype(itype)
low = np.array(estimator.clip_min, dtype=dtype)
high = np.array(estimator.clip_max, dtype=dtype)
# ── Primary output: Clip ──────────────────────────────────────────────────
_clipped = g.op.Clip(X, low, high, name=name, outputs=outputs[:1])
# ── Optional extra output: clip mask ─────────────────────────────────────
if g.convert_options.has("clip_mask", estimator, name):
assert (
len(outputs) > 1
), f"Expected at least 2 outputs when clip_mask is active, got {len(outputs)}"
below = g.op.Less(X, low, name=f"{name}_below")
above = g.op.Greater(X, high, name=f"{name}_above")
g.op.Or(below, above, name=f"{name}_mask", outputs=outputs[1:2])
return outputs[0] if len(outputs) == 1 else tuple(outputs)
4. Training data#
rng = np.random.default_rng(0)
X_train = rng.standard_normal((80, 4)).astype(np.float32)
X_test = rng.standard_normal((20, 4)).astype(np.float32)
transformer = ClipTransformer(clip_min=-0.5, clip_max=0.5).fit(X_train)
5. Baseline conversion — no extra output#
Without any convert_options the model produces a single output.
onx_plain = to_onnx(
transformer, (X_train,), extra_converters={ClipTransformer: convert_clip_transformer}
)
print("=== Plain conversion (no clip_mask) ===")
print(f"Outputs: {[o.name for o in onx_plain.graph.output]}")
sess_plain = onnxruntime.InferenceSession(
onx_plain.SerializeToString(), providers=["CPUExecutionProvider"]
)
(clipped_onnx,) = sess_plain.run(None, {"X": X_test})
clipped_sklearn = transformer.transform(X_test)
assert np.allclose(clipped_sklearn, clipped_onnx, atol=1e-6), "Clipped values differ!"
print("Clipped values match sklearn ✓")
=== Plain conversion (no clip_mask) ===
Outputs: ['Y']
Clipped values match sklearn ✓
6. Conversion with clip_mask=True#
Passing ClipOptions(clip_mask=True) instructs the framework to allocate
a second output slot. The converter detects this via
g.convert_options.has("clip_mask", estimator) and emits the boolean mask.
clip_opts = ClipOptions(clip_mask=True)
onx_with_mask = to_onnx(
transformer,
(X_train,),
extra_converters={ClipTransformer: convert_clip_transformer},
convert_options=clip_opts,
)
print("\n=== Conversion with clip_mask=True ===")
print(f"Outputs: {[o.name for o in onx_with_mask.graph.output]}")
sess_mask = onnxruntime.InferenceSession(
onx_with_mask.SerializeToString(), providers=["CPUExecutionProvider"]
)
clipped_onnx2, mask_onnx = sess_mask.run(None, {"X": X_test})
# Verify clipped values
assert np.allclose(clipped_sklearn, clipped_onnx2, atol=1e-6), "Clipped values differ!"
print("Clipped values match sklearn ✓")
# Verify boolean mask
expected_mask = transformer.get_clip_mask(X_test)
assert np.array_equal(expected_mask, mask_onnx), "Clip mask differs!"
print("Clip mask matches sklearn ✓")
print(f"\nmask_onnx shape : {mask_onnx.shape}")
print(f"fraction clipped : {mask_onnx.mean():.2%}")
=== Conversion with clip_mask=True ===
Outputs: ['Y', 'clip_mask']
Clipped values match sklearn ✓
Clip mask matches sklearn ✓
mask_onnx shape : (20, 4)
fraction clipped : 57.50%
7. Visualize the ONNX graph#
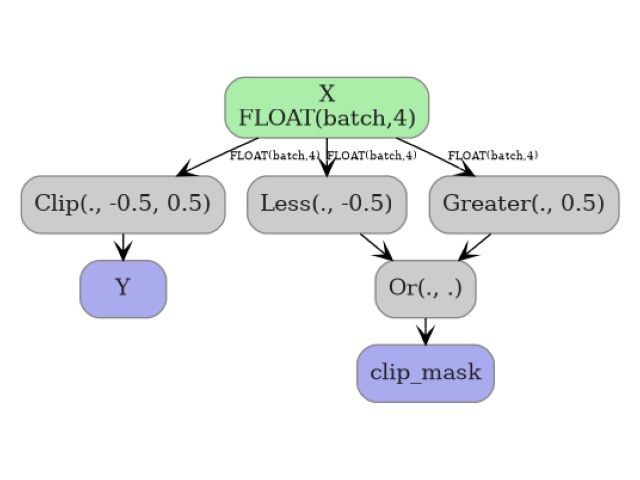
The graph with clip_mask=True contains the Clip node for the primary
output plus Less, Greater, and Or nodes for the mask.
plot_dot(onx_with_mask)
Total running time of the script: (0 minutes 0.326 seconds)
Related examples
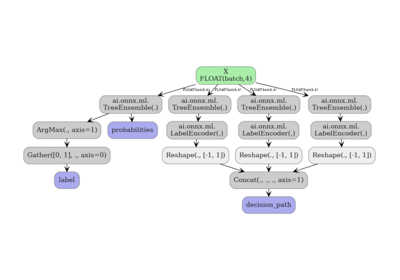
Exporting sklearn tree models with convert options
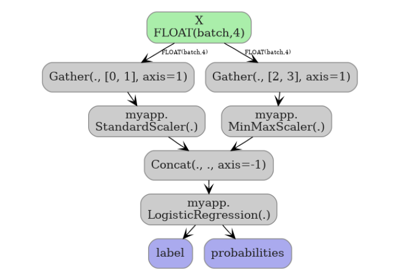
Exporting sklearn estimators as ONNX local functions