Note
Go to the end to download the full example code.
Computed Shapes: Add + Concat + Reshape#
This example shows how BasicShapeBuilder tracks symbolic dimension
expressions through a sequence of Add, Concat, and Reshape nodes,
and compares the result with the standard
onnx.shape_inference.infer_shapes() and the
onnx-shape-inference
package from PyPI.
The key difference is that onnx.shape_inference.infer_shapes can only
propagate shapes when dimensions are statically known integers. When the
model contains dynamic (symbolic) dimensions it typically assigns None
(unknown) to most intermediate results. BasicShapeBuilder instead
keeps the dimensions as symbolic arithmetic expressions so that output shapes
are expressed in terms of the input dimension names.
The onnx-shape-inference package also performs symbolic shape inference
using SymPy to track dimension expressions across
nodes. It operates on the onnx-ir representation of the model.
See ShapeBuilder for a detailed description of how
BasicShapeBuilder
works and a comparison table with onnx.shape_inference.infer_shapes().
import numpy as np
import pandas
import onnx
import onnx_ir as ir
import onnxruntime
import onnx.helper as oh
import onnx.numpy_helper as onh
from onnx_shape_inference import infer_symbolic_shapes
from yobx.xshape import BasicShapeBuilder
TFLOAT = onnx.TensorProto.FLOAT
TINT64 = onnx.TensorProto.INT64
Helper functions for shape inference#
The three shape-inference approaches used throughout this example are each wrapped in a small helper so the same logic can be reused without repetition.
def infer_shapes_onnx(model: onnx.ModelProto) -> dict:
"""Run :func:`onnx.shape_inference.infer_shapes`; return ``{name: shape}``."""
inferred = onnx.shape_inference.infer_shapes(model)
shapes = {}
for vi in [*inferred.graph.input, *inferred.graph.value_info, *inferred.graph.output]:
t = vi.type.tensor_type
if t.HasField("shape"):
shapes[vi.name] = tuple(
d.dim_param if d.dim_param else (d.dim_value if d.dim_value else None)
for d in t.shape.dim
)
else:
shapes[vi.name] = "unknown"
return shapes
def infer_shapes_onnx_ir(model: onnx.ModelProto) -> dict:
"""Run onnx-shape-inference :func:`infer_symbolic_shapes`; return ``{name: shape}``."""
ir_model = ir.serde.deserialize_model(model)
ir_model = infer_symbolic_shapes(ir_model)
shapes = {}
for v in ir_model.graph.inputs:
shapes[v.name] = str(v.shape)
for node in ir_model.graph:
for out in node.outputs:
shapes[out.name] = str(out.shape)
return shapes
def infer_shapes_basic(model: onnx.ModelProto) -> BasicShapeBuilder:
"""Run :class:`BasicShapeBuilder` over *model*; return the populated builder."""
b = BasicShapeBuilder()
b.run_model(model)
return b
def print_shapes(shapes, names: list) -> None:
"""Print shapes for *names* from *shapes*.
*shapes* may be either a ``{name: shape}`` dict (as returned by
:func:`infer_shapes_onnx` and :func:`infer_shapes_onnx_ir`) or a
:class:`BasicShapeBuilder` instance (as returned by
:func:`infer_shapes_basic`).
"""
for name in names:
if isinstance(shapes, dict):
shape = shapes.get(name, "unknown")
else:
shape = shapes.get_shape(name)
print(f" {name:15s} shape={shape}")
def make_shape_comparison_table(model: onnx.ModelProto, names: list) -> pandas.DataFrame:
"""Build a side-by-side shape comparison DataFrame for *model*.
Runs all three inference tools and returns a :class:`pandas.DataFrame`
with one row per tensor name and one column per tool.
Columns: ``onnx``, ``onnx_ir``, ``basic``.
"""
onnx_shapes = infer_shapes_onnx(model)
ir_shapes = infer_shapes_onnx_ir(model)
basic = infer_shapes_basic(model)
rows = []
for name in names:
rows.append(
{
"name": name,
"onnx": str(onnx_shapes.get(name, "unknown")),
"onnx_ir": str(ir_shapes.get(name, "unknown")),
"basic": str(basic.get_shape(name)),
}
)
return pandas.DataFrame(rows).set_index("name")
Build a small model#
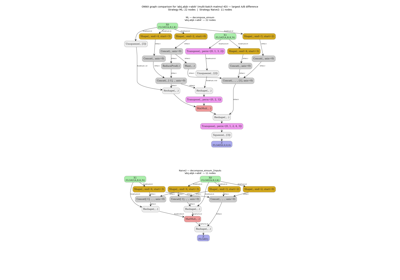
The graph performs the following steps:
Add(X, Y)— element-wise addition of two tensors with shape(batch, seq, d_model).Concat(added, X, axis=2)— concatenate the result with the originalXalong the last axis, giving shape(batch, seq, 2*d_model).Reshape(concat_out, shape)— flatten the last two dimensions using a fixed shape constant[0, 0, -1], which collapses(batch, seq, 2*d_model)back to(batch, seq, 2*d_model).
model = oh.make_model(
oh.make_graph(
[
oh.make_node("Add", ["X", "Y"], ["added"]),
oh.make_node("Concat", ["added", "X"], ["concat_out"], axis=2),
oh.make_node("Reshape", ["concat_out", "reshape_shape"], ["Z"]),
],
"add_concat_reshape",
[
oh.make_tensor_value_info("X", TFLOAT, ["batch", "seq", "d_model"]),
oh.make_tensor_value_info("Y", TFLOAT, ["batch", "seq", "d_model"]),
],
[oh.make_tensor_value_info("Z", TFLOAT, [None, None, None])],
[onh.from_array(np.array([0, 0, -1], dtype=np.int64), name="reshape_shape")],
),
opset_imports=[oh.make_opsetid("", 18)],
ir_version=10,
)
Shape inference with ONNX#
onnx.shape_inference.infer_shapes propagates shapes through the model.
For dynamic dimensions the inferred shapes for intermediate results are
often unknown (None).
=== onnx.shape_inference.infer_shapes ===
X shape=('batch', 'seq', 'd_model')
Y shape=('batch', 'seq', 'd_model')
added shape=('batch', 'seq', 'd_model')
concat_out shape=('batch', 'seq', 'unk__0')
Z shape=('batch', 'seq', 'unk__1')
Shape inference with onnx-shape-inference (PyPI)#
The onnx-shape-inference
package offers a second symbolic approach. It works on the
onnx_ir.Model representation and uses SymPy to track dimension
expressions. Install it with pip install onnx-shape-inference.
Compared with onnx.shape_inference.infer_shapes, it successfully
resolves the Concat output to (batch, seq, 2*d_model). The
Reshape output receives a freshly-generated symbol (_d0) because
the [0, 0, -1] constant shape tensor is not yet fully evaluated by this
library.
onnx_ir_shapes = infer_shapes_onnx_ir(model)
print("=== onnx-shape-inference (infer_symbolic_shapes) ===")
print_shapes(onnx_ir_shapes, ["X", "Y", "added", "concat_out", "Z"])
=== onnx-shape-inference (infer_symbolic_shapes) ===
X shape=[batch,seq,d_model]
Y shape=[batch,seq,d_model]
added shape=[batch,seq,d_model]
concat_out shape=[batch,seq,2*d_model]
Z shape=[batch,seq,_d0]
Shape inference with BasicShapeBuilder#
BasicShapeBuilder
keeps the shapes as symbolic expressions. Because reshape_shape is a
constant [0, 0, -1], the builder can evaluate the Reshape and express
the output shape as a function of the input dimensions.
builder = infer_shapes_basic(model)
print("\n=== BasicShapeBuilder ===")
print_shapes(builder, ["X", "Y", "added", "concat_out", "Z"])
=== BasicShapeBuilder ===
X shape=('batch', 'seq', 'd_model')
Y shape=('batch', 'seq', 'd_model')
added shape=('batch', 'seq', 'd_model')
concat_out shape=('batch', 'seq', '2*d_model')
Z shape=('batch', 'seq', '2*d_model')
Evaluate symbolic shapes with concrete values#
Once the concrete values of the dynamic dimensions are known,
evaluate_shape resolves
each symbolic expression to its actual integer value.
X concrete shape=(2, 5, 8)
Y concrete shape=(2, 5, 8)
added concrete shape=(2, 5, 8)
concat_out concrete shape=(2, 5, 16)
Z concrete shape=(2, 5, 16)
Verify with real data#
Finally, run the model with concrete numpy arrays and confirm that the
shapes predicted by BasicShapeBuilder match the actual output
shapes.
feeds = {
"X": np.random.rand(2, 5, 8).astype(np.float32),
"Y": np.random.rand(2, 5, 8).astype(np.float32),
}
session = onnxruntime.InferenceSession(
model.SerializeToString(), providers=["CPUExecutionProvider"]
)
outputs = session.run(None, feeds)
result = builder.compare_with_true_inputs(feeds, outputs)
print("\n=== shape comparison ===")
data = []
for name, dims in result.items():
obs = dict(result=name)
for i, dim in enumerate(dims):
for c, v in zip(["expression", "expected", "computed"], dim):
data.append(dict(result=name, dimension=i, col=c, value=v))
print(pandas.DataFrame(data).pivot(index=["result", "dimension"], columns="col", values="value"))
=== shape comparison ===
col computed expected expression
result dimension
Z 0 2 2 batch
1 5 5 seq
2 16 16 2*d_model
Impact of named output shapes on constraints (NonZero)#
Some operators—such as NonZero—introduce a fresh symbolic dimension for
their output because the number of results depends on the values of the
input tensor, not merely its shape. BasicShapeBuilder assigns an
internal name like NEWDIM_nonzero_0 to that dimension.
When the graph output is declared without named dimensions (all None),
the internal name is kept as-is and no constraint is registered.
When the graph output is declared with named dimensions (e.g.
["rank", "nnz"]), run_value_info detects
the mismatch between the computed internal name NEWDIM_nonzero_0 and the
user-supplied name nnz, and registers the constraint
NEWDIM_nonzero_0 = nnz. The dimension naming step then renames the
internal token to the user-visible name across all shapes.
The two models share the same 7-node graph topology:
Abs → Relu → Add → Mul → NonZero(nz) → Transpose → Cast(nz_float).
They differ only in how the graph outputs are annotated.
_NZ_NODES = [
oh.make_node("Abs", ["X"], ["abs_out"]),
oh.make_node("Relu", ["abs_out"], ["relu_out"]),
oh.make_node("Add", ["relu_out", "relu_out"], ["double_out"]),
oh.make_node("Mul", ["double_out", "relu_out"], ["mul_out"]),
oh.make_node("NonZero", ["mul_out"], ["nz"]),
oh.make_node("Transpose", ["nz"], ["transposed_nz"]),
oh.make_node("Cast", ["transposed_nz"], ["nz_float"], to=TFLOAT),
]
_NZ_INPUT = [oh.make_tensor_value_info("X", TFLOAT, ["batch", "seq"])]
_NZ_NAMES = [
"X",
"abs_out",
"relu_out",
"double_out",
"mul_out",
"nz",
"transposed_nz",
"nz_float",
]
Anonymous output shapes ([None, None]): the data-dependent dimension
keeps the internal placeholder NEWDIM_nonzero_0 and no constraint is
registered.
nz_model_anon = oh.make_model(
oh.make_graph(
_NZ_NODES,
"nonzero_anon",
_NZ_INPUT,
[
oh.make_tensor_value_info("nz", TINT64, [None, None]),
oh.make_tensor_value_info("nz_float", TFLOAT, [None, None]),
],
),
opset_imports=[oh.make_opsetid("", 18)],
ir_version=10,
)
Named output shapes (["rank", "nnz"]): the constraint
NEWDIM_nonzero_0 = nnz is registered and the placeholder is renamed
throughout the graph.
nz_model_named = oh.make_model(
oh.make_graph(
_NZ_NODES,
"nonzero_named",
_NZ_INPUT,
[
oh.make_tensor_value_info("nz", TINT64, ["rank", "nnz"]),
oh.make_tensor_value_info("nz_float", TFLOAT, ["do1", "do2"]),
],
),
opset_imports=[oh.make_opsetid("", 18)],
ir_version=10,
)
Comparison table — anonymous output shapes#
With [None, None] output annotations the data-dependent dimension is
kept as the internal placeholder NEWDIM_nonzero_0 by
BasicShapeBuilder; no constraint is registered.
print("=== anonymous output shapes ===")
print(make_shape_comparison_table(nz_model_anon, _NZ_NAMES).to_string())
=== anonymous output shapes ===
onnx onnx_ir basic
name
X ('batch', 'seq') [batch,seq] ('batch', 'seq')
abs_out ('batch', 'seq') [batch,seq] ('batch', 'seq')
relu_out ('batch', 'seq') [batch,seq] ('batch', 'seq')
double_out ('batch', 'seq') [batch,seq] ('batch', 'seq')
mul_out ('batch', 'seq') [batch,seq] ('batch', 'seq')
nz (2, 'unk__0') [2,_d0] (2, 'NEWDIM_nonzero_0')
transposed_nz ('unk__0', 2) [_d0,2] ('NEWDIM_nonzero_0', 2)
nz_float ('unk__0', 2) [_d0,2] ('NEWDIM_nonzero_0', 2)
Registered constraints (anonymous model):
anon_builder = infer_shapes_basic(nz_model_anon)
print("constraints:", anon_builder.get_registered_constraints())
constraints: {}
Comparison table — named output shapes#
With ["rank", "nnz"] output annotations BasicShapeBuilder
registers the constraint NEWDIM_nonzero_0 = nnz and renames the
placeholder everywhere, so nz shape becomes (2, 'nnz') and the
propagation continues through Transpose and Cast.
print("=== named output shapes ===")
print(make_shape_comparison_table(nz_model_named, _NZ_NAMES).to_string())
=== named output shapes ===
onnx onnx_ir basic
name
X ('batch', 'seq') [batch,seq] ('batch', 'seq')
abs_out ('batch', 'seq') [batch,seq] ('batch', 'seq')
relu_out ('batch', 'seq') [batch,seq] ('batch', 'seq')
double_out ('batch', 'seq') [batch,seq] ('batch', 'seq')
mul_out ('batch', 'seq') [batch,seq] ('batch', 'seq')
nz (2, 'nnz') [2,nnz] (2, 'do1')
transposed_nz ('nnz', 2) [nnz,2] ('do1', 2)
nz_float ('do1', 2) [do1,2] ('do1', 2)
Registered constraints (named model):
named_builder = infer_shapes_basic(nz_model_named)
print("constraints:", named_builder.get_registered_constraints())
constraints: {'NEWDIM_nonzero_0': {'nnz', 'do1'}}
Total running time of the script: (0 minutes 0.409 seconds)
Related examples

Computation Cost: How It Works and Supported Operator Formulas
Comparing Computational Cost of Three Einsum→ONNX Strategies
