Note
Go to the end to download the full example code.
MiniOnnxBuilder: serialize tensors to an ONNX model#
MiniOnnxBuilder
creates minimal ONNX models whose only purpose is to store tensors as
initializers and return them when the model is executed. The model has
no inputs — running it simply replays the stored values.
This is useful for:
capturing intermediate activations or model weights for debugging,
persisting arbitrary nested Python structures (dicts, tuples, lists, torch tensors,
DynamicCache…) in a standard, portable format,sharing small test fixtures without committing raw binary files.
The module also provides two higher-level helpers built on top of
MiniOnnxBuilder:
create_onnx_model_from_input_tensors— serialize any nested structure to anonnx.ModelProto.create_input_tensors_from_onnx_model— deserialize the model back to the original Python structure.
import numpy as np
import onnxruntime
import torch
from yobx.helpers.mini_onnx_builder import (
MiniOnnxBuilder,
create_onnx_model_from_input_tensors,
create_input_tensors_from_onnx_model,
)
1. Store a single numpy array#
The simplest use-case: add one initializer as an output and recover it.
builder = MiniOnnxBuilder()
weights = np.array([1.0, 2.0, 3.0], dtype=np.float32)
builder.append_output_initializer("weights", weights)
model = builder.to_onnx()
ref = onnxruntime.InferenceSession(model.SerializeToString(), providers=["CPUExecutionProvider"])
(recovered,) = ref.run(None, {})
print("original :", weights)
print("recovered:", recovered)
assert np.array_equal(weights, recovered)
original : [1. 2. 3.]
recovered: [1. 2. 3.]
2. Store multiple tensors (numpy + torch)#
Several calls to append_output_initializer
add more outputs to the same model.
builder2 = MiniOnnxBuilder()
x_np = np.arange(6, dtype=np.int64).reshape(2, 3)
x_torch = torch.tensor([[0.1, 0.2], [0.3, 0.4]], dtype=torch.float32)
builder2.append_output_initializer("x_np", x_np)
builder2.append_output_initializer("x_torch", x_torch.numpy())
model2 = builder2.to_onnx()
ref2 = onnxruntime.InferenceSession(
model2.SerializeToString(), providers=["CPUExecutionProvider"]
)
got_np, got_torch = ref2.run(None, {})
print("x_np :", got_np)
print("x_torch:", got_torch)
assert np.array_equal(x_np, got_np)
assert np.allclose(x_torch.numpy(), got_torch)
x_np : [[0 1 2]
[3 4 5]]
x_torch: [[0.1 0.2]
[0.3 0.4]]
3. Store a sequence of tensors#
append_output_sequence
wraps multiple tensors into an ONNX Sequence.
builder3 = MiniOnnxBuilder()
seq = [np.array([10, 20], dtype=np.int64), np.array([30, 40], dtype=np.int64)]
builder3.append_output_sequence("my_seq", seq)
model3 = builder3.to_onnx()
ref3 = onnxruntime.InferenceSession(
model3.SerializeToString(), providers=["CPUExecutionProvider"]
)
(got_seq,) = ref3.run(None, {})
print("sequence:", got_seq)
for original, restored in zip(seq, got_seq):
assert np.array_equal(original, restored)
sequence: [array([10, 20], dtype=int64), array([30, 40], dtype=int64)]
4. Round-trip a nested Python structure#
The higher-level helpers handle arbitrary nesting of dicts, tuples, lists, numpy arrays and torch tensors automatically.
inputs = {
"ids": np.array([1, 2, 3], dtype=np.int64),
"mask": np.array([1, 1, 0], dtype=np.bool_),
"hidden": torch.randn(2, 4, dtype=torch.float32),
}
proto = create_onnx_model_from_input_tensors(inputs)
restored = create_input_tensors_from_onnx_model(proto)
print("keys:", list(restored.keys()))
for k in inputs:
print(f" {k}: {inputs[k].shape} -> {restored[k].shape}")
if isinstance(inputs[k], np.ndarray):
assert np.array_equal(inputs[k], restored[k]), f"mismatch for {k}"
else:
assert torch.equal(inputs[k], restored[k]), f"mismatch for {k}"
keys: ['ids', 'mask', 'hidden']
ids: (3,) -> (3,)
mask: (3,) -> (3,)
hidden: torch.Size([2, 4]) -> torch.Size([2, 4])
5. Randomize float tensors to save space#
When randomize=True the actual weight values are replaced by a
random-number generator node, keeping the shape and dtype but
discarding the original values. This drastically reduces model size
for large weight tensors when exact values are not needed.
big = np.random.randn(128, 256).astype(np.float32)
proto_rand = create_onnx_model_from_input_tensors(big, randomize=True)
proto_exact = create_onnx_model_from_input_tensors(big)
print(f"randomized model size : {proto_rand.ByteSize():>8} bytes")
print(f"exact model size : {proto_exact.ByteSize():>8} bytes")
assert proto_rand.ByteSize() < proto_exact.ByteSize()
randomized model size : 225 bytes
exact model size : 131247 bytes
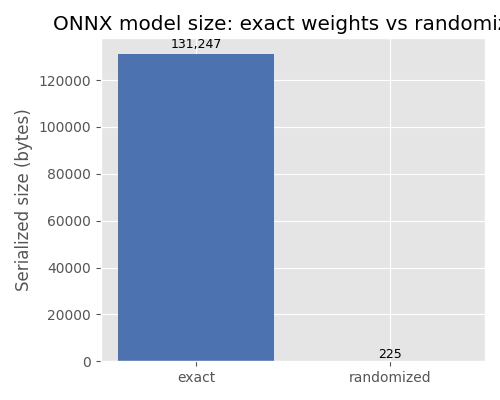
Plot: model size comparison#
The bar chart below illustrates the difference in serialized model size
between a model that stores the actual weight values (exact) and one
that replaces them with a random-number generator node (randomized).
import matplotlib.pyplot as plt # noqa: E402
sizes = [proto_exact.ByteSize(), proto_rand.ByteSize()]
labels = ["exact", "randomized"]
fig, ax = plt.subplots(figsize=(5, 4))
bars = ax.bar(labels, sizes, color=["#4c72b0", "#dd8452"])
ax.set_ylabel("Serialized size (bytes)")
ax.set_title("ONNX model size: exact weights vs randomized")
for bar, size in zip(bars, sizes):
ax.text(
bar.get_x() + bar.get_width() / 2,
bar.get_height() * 1.01,
f"{size:,}",
ha="center",
va="bottom",
fontsize=9,
)
plt.tight_layout()
plt.show()
Total running time of the script: (0 minutes 0.156 seconds)
Related examples
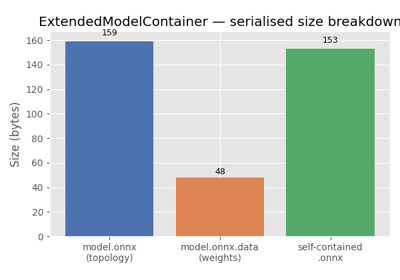
ExtendedModelContainer: large-initializer ONNX models