Note
Go to the end to download the full example code.
Using sklearn-onnx to convert any scikit-learn estimator#
yobx.sklearn.to_onnx() ships with built-in converters for a curated
set of scikit-learn estimators (StandardScaler,
LogisticRegression, DecisionTree*, MLP*, and Pipeline).
For any estimator that is not covered by the built-in registry, you can
supply your own converter through the extra_converters keyword argument.
wrap_skl2onnx_converter() makes the conversion from
sklearn-onnx. One line of setup after
looking up the skl2onnx converter function.
import numpy as np
import onnxruntime
from sklearn.neural_network import MLPClassifier
from yobx import doc
from yobx.sklearn import to_onnx, wrap_skl2onnx_converter
Converter made for skl2onnx#
We take it from sklearn-onnx.
import skl2onnx
convert_sklearn_mlp_classifier = (
skl2onnx.operator_converters.multilayer_perceptron.convert_sklearn_mlp_classifier
)
Convert to ONNX using the factory helper#
wrap_skl2onnx_converter() makes the function look
like a converter for yobx.`
X = np.array([[1, 2], [3, 4], [5, 6], [7, 8]], dtype=np.float32)
y = np.array([0, 0, 1, 1])
mlp = MLPClassifier(hidden_layer_sizes=(4,), activation="relu", max_iter=2000)
mlp.fit(X, y)
converter = wrap_skl2onnx_converter(convert_sklearn_mlp_classifier)
artifact = to_onnx(mlp, (X,), extra_converters={MLPClassifier: converter})
ref_b = onnxruntime.InferenceSession(
artifact.SerializeToString(), providers=["CPUExecutionProvider"]
)
results_b = ref_b.run(None, {"X": X})
label_b, proba_b = results_b[0], results_b[1]
np.testing.assert_array_equal(mlp.predict(X), label_b)
np.testing.assert_allclose(mlp.predict_proba(X), proba_b, atol=1e-5)
print("Predictions match ✓")
~/vv/this312/lib/python3.12/site-packages/sklearn/neural_network/_multilayer_perceptron.py:785: ConvergenceWarning: Stochastic Optimizer: Maximum iterations (2000) reached and the optimization hasn't converged yet.
warnings.warn(
Predictions match ✓
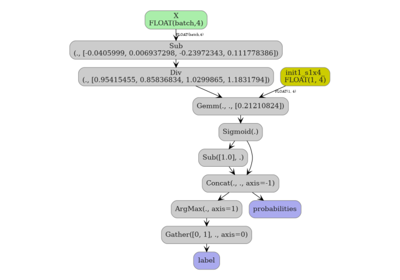
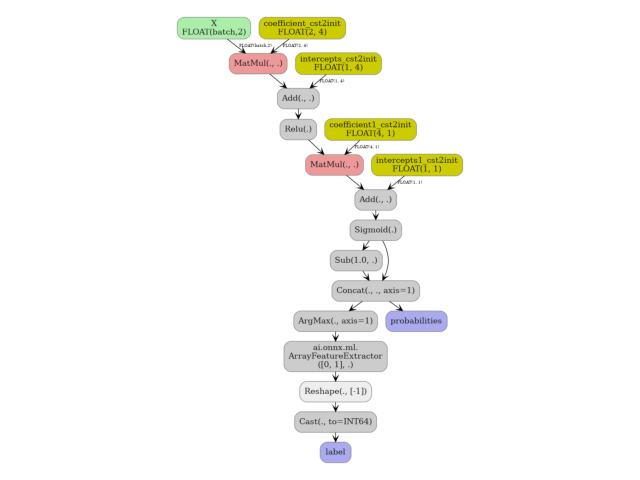
4. Visualise the exported ONNX graph#
yobx.doc.plot_dot() renders the onnx.ModelProto as a
directed graph using Graphviz. The nodes emitted by the skl2onnx
converter are injected directly into the enclosing graph.
doc.plot_dot(artifact)
Total running time of the script: (0 minutes 0.666 seconds)
Related examples
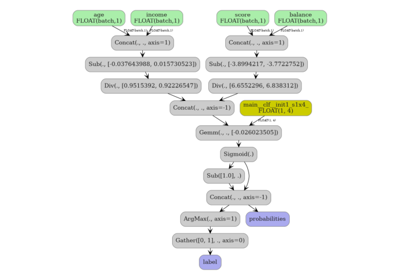
DataFrame input to a Pipeline with ColumnTransformer

Float32 vs Float64: precision loss with PLSRegression